Efficiently Unpickling from Buffer Objects
I learned to use the python psycopg2 module in a hurry a while back. I hit a stumbling block: I couldn't figure out how to populate bytea columns. It turns out to be much simpler, you just use a psycopg2.Binary object in the call to the execute statement on a cursor. However, you never get back a psycopg2.Binary from cursors. Instead, you get back a buffer object. In my case I am pickling Python dictionaries and there is no pickle function that will directly unpickle from a buffer object. The StringIO module does wrap the buffer object allowing the pickle.load method to read from the buffer. Here is some example code showing how I did it.
import psycopg2 import pickle import StringIO conn = psycopg2.connect(user='postgres',database='postgres') cur = conn.cursor() cur.execute('Select %s', (psycopg2.Binary(pickle.dumps({'foo':'bar'},-1)), )) result, = cur.fetchone() print type(result) cur.close() conn.rollback() result = StringIO.StringIO(result) print pickle.load(result)
This code constructs a psycopg2.Binary from a string returned by pickle.dumps. The pickled object is a plain python dictionary. The object is passed to a SQL SELECT statement which promptly returns it. Running this gives a result of
<type 'buffer'>
{'foo': 'bar'}
I call type on the result to show that it indeed is a buffer object. It unpickles just fine using StringIO.StringIO.
Why store a blob in the database?
Before I go any farther, I'd like to caution you against storing binary data in your relational database. There are plenty of reasons to avoid it. First off, you can't really do any meaningful queries on a blob. PostgreSQL actually supports a native key value column type called hstore. When you create an hstore column, each row in a table can store arbitrary data and you can query against it. A key-value store cannot handle arbitrarily nested objects however. PostgreSQL also supports operations on JSON natively. JSON of course can handle arbitrarily nested objects. However, you can't store binary data in JSON. You'd need to escape it using something like base64 or base16. At that point, you've greatly inflated the size of your data.
Before storing blobs in a relational database, be sure to consider all your other options.
Finding the fastest way out of a pickle
The need to unpickle buffer objects is interesting because it uses both the pickle and StringIO modules. Both of these modules have pure-C implementations with slightly less functionality. They are cPickle and cStringIO respectively. I was curious to see what combination of the modules was the fastest. All code and benchmarks performed here are done using Stackless Python 2.7.5.
Preliminary Investigation
The first question I had was how much copying of data was going on behind the scenes when StringIO.StringIO(buffer) is executed. Looking into the implementation of the StringIO module shows the following in Lib/StringIO.py
# Force self.buf to be a string or unicode if not isinstance(buf, basestring): buf = str(buf)
The argument buf is the argument of the constructor StringIO.StringIO. If what you pass it is is not a string, it gets converted to one. A deep copy is made when str is called.
Looking at the cStringIO module I found out that cStringIO.StringIO is actually a factory function. If you pass it a parameter, you get a cStringIO.StringI object back. The constructor for that object is shown here
static PyObject * newIobject(PyObject *s) { Iobject *self; Py_buffer buf; PyObject *args; int result; args = Py_BuildValue("(O)", s); if (args == NULL) return NULL; result = PyArg_ParseTuple(args, "s*:StringIO", &buf); Py_DECREF(args); if (!result) return NULL; self = PyObject_New(Iobject, &Itype); if (!self) { PyBuffer_Release(&buf); return NULL; } self->buf=buf.buf; self->string_size=buf.len; self->pbuf=buf; self->pos=0; return (PyObject*)self; }
The parameter to cStringIO.StringIO is extracted using PyArg_ParseTuple. The meaning of s* extracts a pointer to any buffer compatible object. Since this doesn't create a copy of the object, it means that the cStringIO module should have a huge performance advantage.
Benchmarking
Since I'm concerned with the case of retrieving a pickled data structure from PostgreSQL, I'm only interested to see how performance improves on dictionaries that contain dictionaries. If all that is being stored is strings, integers, etc. it would not make any sense to pickle them before storing them in the database.
I decided to use the timeit module to measure the performance of just unpickling the data from a buffer object. It's very rare that I write to the database, so the performance of pickling the object is less consequential. I benchmarked all four combinations of the modules. The computer I benchmarked on has CPU-throttling disabled in the BIOS. It's an older Core i7 running Ubuntu 12.04 LTS with all the latest updates. Stackless Python was compiled myself from the source tarball with the default options.
Results
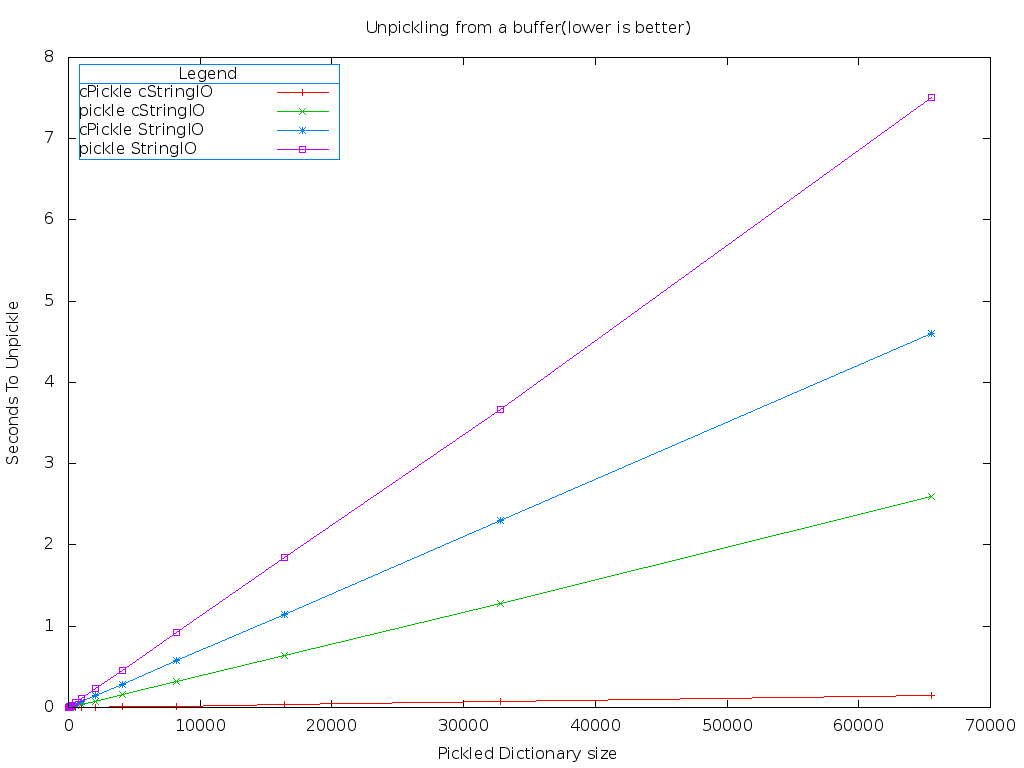
It turns, using cStringIO and cPickle is much, much faster than the default implementations.
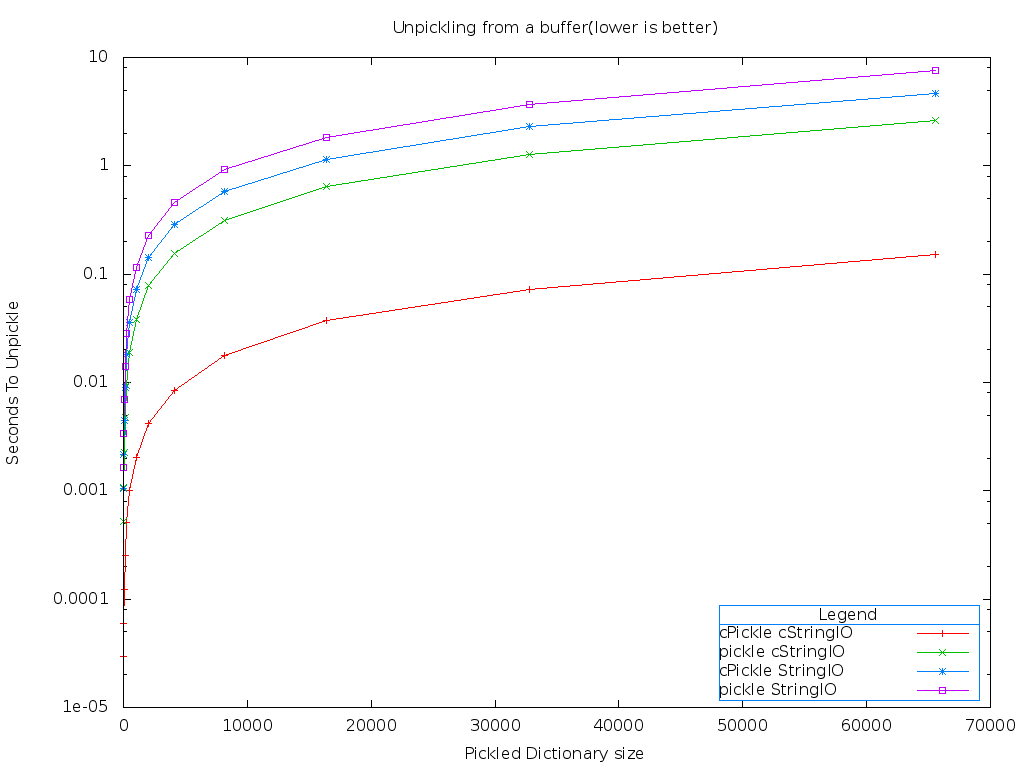
That red line down on the bottom is the combination of the cPickle and cStringIO modules. Lower is better on this benchmark. It's much easier to see the differences using a logarithmic graph.
As expected, all of the functions scale linearly. But using cPickle and cStringIO is over 40 times faster than the pickle and StringIO modules!
How does PyPy compare?
A while back I got interested in PyPy. If you can get all the modules you use or alternatives to them working, it sounds very tempting. For my purposes, I've yet to see any cases where PyPy actually speeds up anything. This benchmark sounded like a great opportunity to see how PyPy compared. I built PyPy 2.7.5 with JIT enabled.
Before we jump to the results it's worth mentioning that PyPy has the cStringIO module as a builtin. It is implemented in RPython, meaning it should be optimized by the JIT compiler that is part of PyPy. The implementation of cPickle on PyPy has the following header
# # Reimplementation of cPickle, mostly as a copy of pickle.py #
So cPickle is not a native-code module on PyPy by any means.
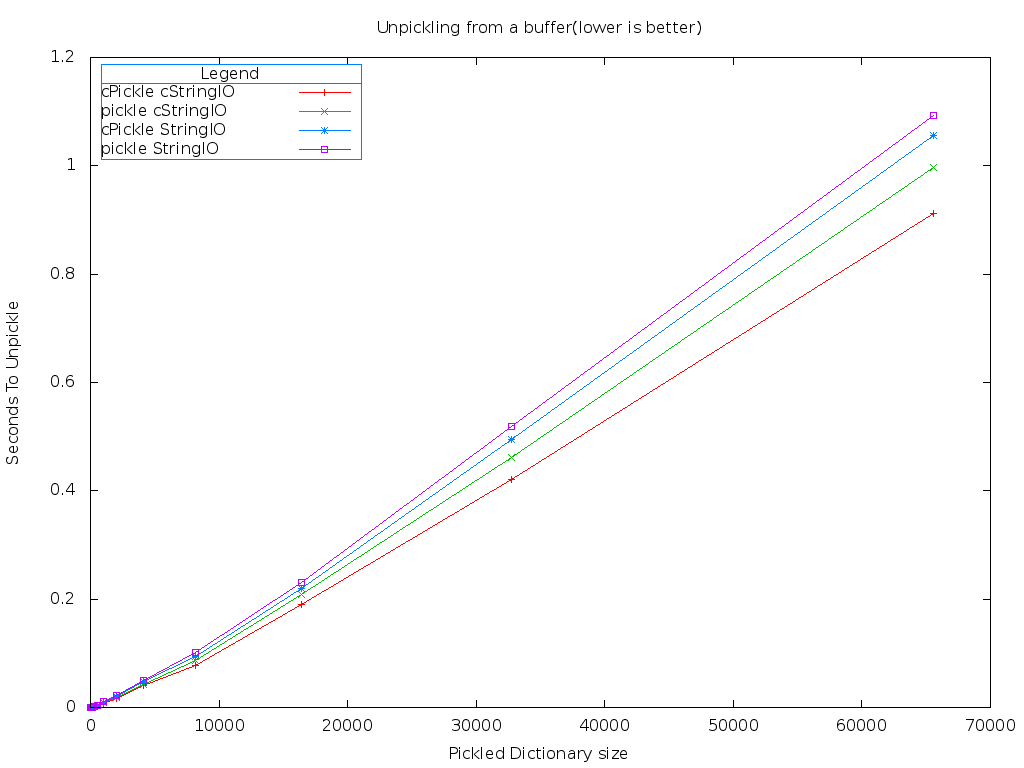
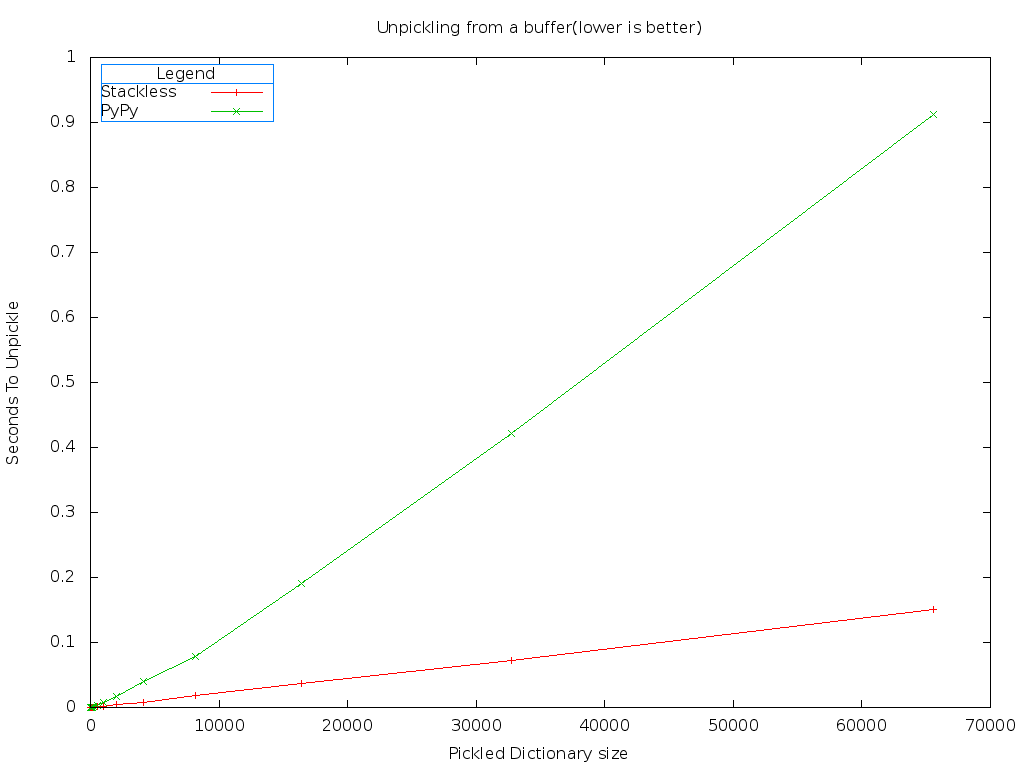
One obvious difference is there isn't any significant gap between the combinations of modules on PyPy. The fastest combination of modules when using PyPy is still cStringIO and cPickle. Let's compare the best case of Stackless against the best case of PyPy.
Stackless is still much faster than PyPy. But worse, PyPy shows slight non-linear behavior at first.
Source Code
The source code along with the data sets from my results is available on github. If enough people run the benchmark and send me the datasets, I'll gladly do a follow up post.